# Trajectories on PCA
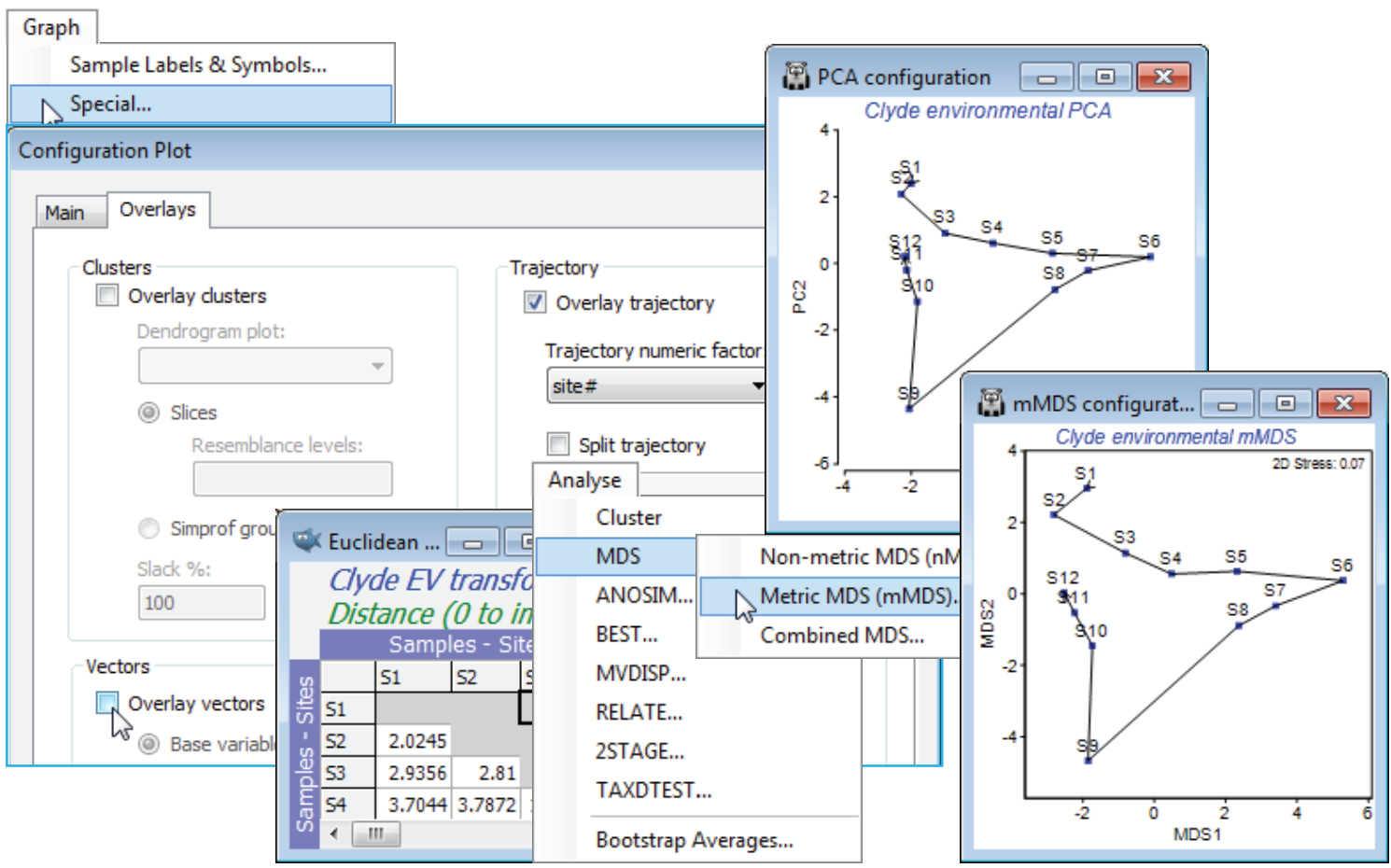
From the **Graph>Special** menu, remove the vector overlay by unchecking the (✓Overlay vectors) box on the **Overlays** tab, and on the same tab, join the points along the transect with (✓Overlay trajectory>Trajectory numeric factor: Site#) – if the factor doesn’t exist, create or import it, as seen under that Ranked variables heading. A better comparison would be of the current PCA with *m*MDS not on the ranked variables but on the same Euclidean distances as created from normalised and transformed variables here, so you may wish to run that **Analyse>MDS>Metric MDS (mMDS)** routine. This indicates one rather obvious difference: *m*MDS works from the resemblance matrix and PCA from the data matrix underlying that. A more important distinction is that *m*MDS does not project the points from the high-d to low-d space as in a PCA, but more carefully arranges them in order optimally to match the low-d Euclidean distance structure to the original distance matrix. Here however, all these ordination cases are effectively indistinguishable: the samples largely lie on a 2-d plane in the 11-d space making it easy for both methods to display an accurate 2-d picture.
[](https://learninghub.primer-e.com/uploads/images/gallery/2024-09/screenshotpage228a.png)
More interesting is the fact that the PCA (or *m*MDS) of the abiotic variables is an excellent match to the *n*MDS of the assemblage (also in Section [11](https://learninghub.primer-e.com/books/primer-v7-user-manual-tutorial/chapter/11-general-data-manipulation-tools-further-pre-treatment)), whether based on biomass, abundance or both, and this observation motivates the BEST routine of Section [13](https://learninghub.primer-e.com/books/primer-v7-user-manual-tutorial/chapter/13-linking-assemblage-to-environment-best-bio-env-linktree). (Note that a PCA of the biota is poor by comparison, since it implicitly uses Euclidean distance rather than an assemblage-based coefficient such as Bray-Curtis – and it actually fails to display a convincing species gradient even though there patently is one there. Choice of a relevant similarity is much the most crucial decision to make in multivariate analysis – a point seen again in Section [14](https://learninghub.primer-e.com/books/primer-v7-user-manual-tutorial/chapter/14-further-matching-of-multivariate-patterns-relate-2stage-best-mvdisp).)