11.2 Example: Plankton (revisited)
We shall show the utility of being able to construct a residual distance matrix and, from this, a residual ordination plot, by reference to a study of differential catches in plankton nets by Winsor & Clarke (1940) , provided by Snedecor (1946) . We met this dataset earlier, as an example of the use of the Wilcoxon signed-rank test. Recall that these data consist of the total log abundance values for each of five different types of plankton (hence, five variables, named using Roman numerals I, II, III, IV and V) that were caught in each of 2 nets towed simultaneously at 2 different depths: one at 29 m and the other at 31 m. There were ten hauls done in this way. The factors associated with these data are:
- Position (either the upper (U) or the lower (L) net: a fixed factor); and
- Haul (10 hauls labeled 1-10: a random factor).
Earlier (in the context of the Wilcoxon signed-rank test) we considered only a single variable - the total sum of the log abundances across all plankton types. Here, however, we are interested in examining the full multivariate dataset with all original $p$ = 5 variables. These data are located in the file ‘Woods_Hole_zooplankton.pri’, found inside the 'Examples_P8 > Woods_Hole_zooplankton' folder.
Visualise the main patterns using ordination
- Open up the file (‘Woods_Hole_zooplankton.pri’) in PRIMER. The data values provided here are already expressed as log abundance values, so there is no need to apply any transformation.
- From the 'Woods_Hole_zooplankton' data sheet, calculate a Bray-Curtis resemblance matrix by clicking Analyse > Resemblance and taking all the defaults (click 'OK'). The result will be 'Resem1', as shown below:
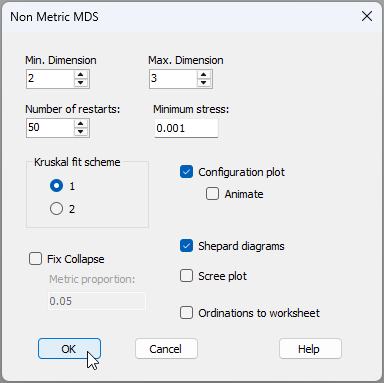
- Next, create a non-metric MDS plot to visualise the patterns of inter-sample relationships among the plankton communities caught in these nets, based on the Bray-Curtis resemblances. From 'Resem1', click Analyse > MDS > Nonmetric MDS (nMDS).... Accept all of the defaults in the 'Non Metric MDS' dialog and just click 'OK'.
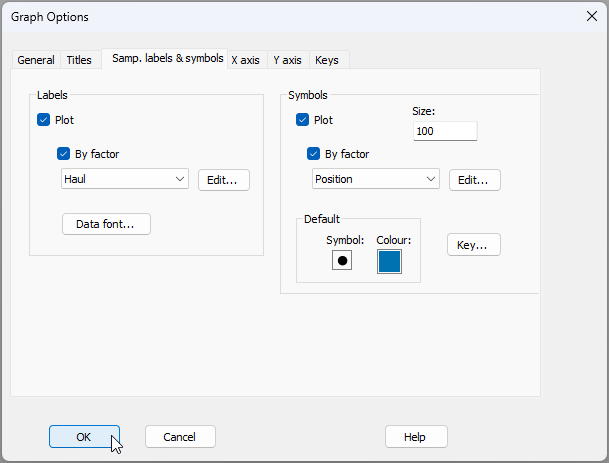
- When you get the multi-plot output, you will see that 'Graph1' in the Explorer tree has the lowest-stress 2D nMDS plot achieved by the routine. From 'Graph1', click Graph > Sample Labels & Symbols... and choose the Labels to be plotted according to the factor of 'Haul', and the Symbols to be plotted according to the factor of 'Position', like so:
The resulting nMDS plot looks like this:
In the above ordination, there do not appear to be any obvious effects on these plankton communities attributable to the factor of 'Position'. Specifically, the symbols corresponding to assemblages caught in Upper ('U') vs Lower ('L') nets are scattered throughout the plot and look to be well mixed throughout this 2D MDS space. If anything, individual 'Hauls' seem to be more important in explaining variation here; we can see assemblages associatd with specific hauls (i.e., the two matching numbers, such as the two 7's, the two 3's, the two 8's, etc.) are often fairly close to one another in this nMDS plot.
However, it is possible that differences between the assemblages from one haul to the next are masking genuine differences due to Position, which may be smaller in size and/or may occur in a different direction than is captured by the nMDS ordination plot of reduced dimension. Of course, we should rely upon a two-way PERMANOVA (or ANOSIM)‡ to formally test for significant effects in either of these factors.¶
Run a two-way PERMANOVA analysis
- Create a design file in accordance with the desired two-factor PERMANOVA model. From 'Resem1', click PERMANOVA+ > Create PERMANOVA Design.... In the resulting design file, called 'Design1', add a row (so there are 2 rows in total), then click inside the appropriate cells in the first column to choose the two factors: 'Haul' and 'Position' in rows 1 and 2, respectively. Also specify the factor of 'Haul' as 'Random' (under 'Type' in column 3), while 'Position' will remain as 'Fixed'. The resulting design file should look like this:
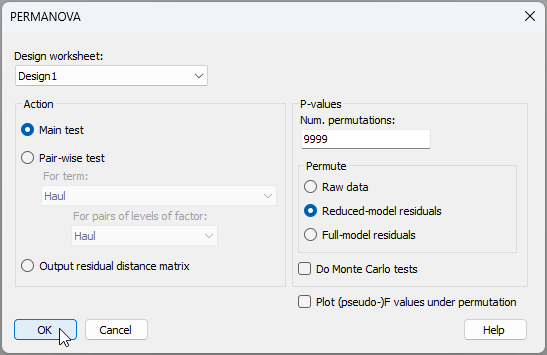
- Now run the PERMANOVA analysis on the basis of this design. From 'Resem1', click PERMANOVA+ > PERMANOVA, then click 'OK' to take all the defaults here (the 'Design worksheet' should default to 'Design1' and all else is fine).
The key results in the output file (called 'PERMANOVA1' in the Explorer tree) look like this:
There is significant variation in these plankton assemblages from haul to haul ($F_{9,9}$ = 6.6, $P$ < 0.001). There is also a significant effect caused by the position (relative depth) of the nets ($F_{1,9}$ = 5.9, $P$ < 0.05). The effect size for the factor of 'Position' was much smaller than that for 'Haul' (see the relative sizes of these in the table labeled 'Estimates of components of variation' in the above output). We can therefore conclude that, in the initial nMDS plot, the significant effects due to the position of the nets are effectively being masked by substantial variation among hauls.
It would be good to remove the effects of different hauls so we can visualise effects due to Position. To do this, we need to:
- Fit a PERMANOVA model with one factor only: 'Haul', and choose the option to output a residual distance matrix.
- Do a non-metric MDS of the residual distance matrix (with the same symbols as above) to examine patterns with respect to variation in the remaining factor: 'Position'.
Note that, if the PERMANOVA analysis had been done and no significant effect of 'Position' had been detected, then there would be no particular reason to pursue the idea of 'digging deeper' in an attempt to visualise its effects.
Obtain residual distances and a residual ordination plot
- To 'remove' the haul effects, we need to fit a one-way PERMANOVA model with the factor of 'Haul' alone. You could just modify the existing design file, but let's make a new design file, for clarity. From 'Resem1', click PERMANOVA+ > Create PERMANOVA Design..., and make the resulting design file (called 'Design2') have just one factor (i.e., 'Haul', the thing we are trying to remove), so that it looks like this:†
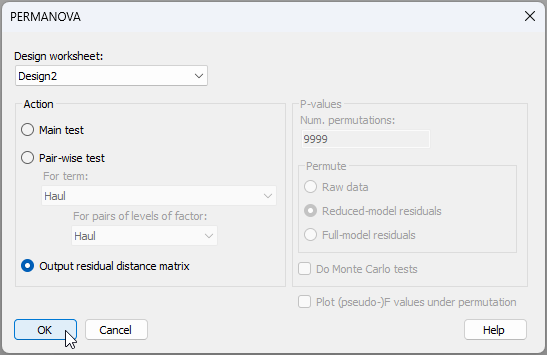
- Let's get the residual distance matrix. From 'Resem1', click PERMANOVA+ > PERMANOVA, choose the 'Design worksheet' as 'Design2', then choose to '$\bullet$Output residual distance matrix', like so:
Notice here, in passing, that the right-hand side of the dialog is 'greyed out' when we choose the option to output a residual distance matrix, as no permutations or tests need to be done in that case. Click 'OK'.
The residual distance matrix is output as 'Resem2' in the Explorer tree, as shown below:
- Finally, we are ready to take a look at our multivariate residuals in an ordination diagram. From 'Resem2', click Analyse > MDS > Nonmetric MDS (nMDS).... Accept all of the defaults in the 'Non Metric MDS' dialog and just click 'OK'. Once the default plot has been produced, click Graph > Sample Labels & Symbols... and choose the Labels to be plotted according to the factor of 'Haul', and the Symbols to be plotted according to the factor of 'Position', just as we had done before.
The resulting lowest-stress 2D nMDS solution for our residual ordination plot (having removed effects due to 'Haul') looks like this:
This is very different from the plot we saw before! Now the effects due to the 'Position' are abundantly clear! This might seem like a spot of magic, but if you sneak a peek back at the original nMDS plot (above), you might notice that, for most of the pairs of observations within a given haul, the 'U' (blue) symbol occurs to the left of the 'L' (amber) symbol. Thus, when we center all of the haul pairs (1 through 10) onto a common centroid, the blue symbols appear to the left and the amber symbols appear to the right. Of course, don't forget that the centering (i.e., the calculation of residual distances) is done in the full-dimensional space, and not in the 2d nMDS space. Being able to carefully work out the patterns we see in a residual plot for a given factor by reference to the original plot in this way is not always possible, however. In fact, it will rarely be the case, as typically the effects of factors are happening across a lot more than 2 dimensions. Nevertheless, this example definitely shows us this fantastic tool for visualising minor effects after removing the effects of some other factor(s) or nuisance variable(s).
‡In this particular example, however, ANOSIM cannot be used for the test of Position for the two-way crossed design, because there is no replication within the cells of the design (i.e., within the Position-by-Haul combinations) and the Position factor has fewer than 3 groups.
¶This is a case of a two-way unreplicated ANOVA design. The lack of replicate nets at the same depth for each haul means that we will be unable to partition out and estimate any potential Position$\times$Haul interactive effects from the estimated Residual variation, with which it is confounded. Despite this inability to separately test for an interaction, it is still important, nevertheless, to fit a two-way design here that includes the 'Haul' factor, and not to fit a one-way model with only the 'Position' factor alone. Specifically, if we ignore the paired nature of the nets in the sampling protocol, we risk being unable to detect 'Position' effects at all. See the section on unreplicated designs in the original PERMANOVA+ manual for details.
†In this particular case (i.e., a one-way PERMANOVA), it does not matter whether we specify 'Haul' as being fixed or random; the residuals will be the same either way. We specify it as random here for clarity and consistency in the specification of our reduced (one-factor) model.

![04.Plankton_net_data_v2[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/04-plankton-net-data-v2-i.png)
![05.Plankton_net_resem_v2[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/05-plankton-net-resem-v2-i.png)
![06.nMDS_Plankton_v2[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/06-nmds-plankton-v2-i.png)
![07.PERMANOVA_design_file_Plankton_v2[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/07-permanova-design-file-plankton-v2-i.png)
![07.PERMANOVA_output_file_Plankton[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/07-permanova-output-file-plankton-i.png)
![08.Design_file_Hauls_only_v2[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/08-design-file-hauls-only-v2-i.png)
![10.Residual_distance_matrix_plankton_v2[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/10-residual-distance-matrix-plankton-v2-i.png)
![11.Residual_ordination_nMDS_Plankton_v2[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/11-residual-ordination-nmds-plankton-v2-i.png)