15.9 New diagnostics for PCA/PCO plots
Background
Consider a cloud of $N$ points (sampling units) in a $p$-dimensional multivariate space. Principal components analysis (PCA) is an ordination method that will find an axis through that cloud of points so as to maximise the total variance of the points, when they are projected at right angles onto that axis. Having found such an axis, a second axis is then built in a similar way, subject to it being perpendicular to (independent of) the first axis, and so on. PCA does this in Euclidean space. Principal coordinate analysis (abbreviated as PCO or PCoA) also does this, but in the space of a chosen resemblance measure ( Gower (1966) ).
Hence, we may conceptually consider either PCA or PCO as a type of projection of the points into a smaller number of dimensions that can capture important major stuctures occurring across the data cloud as a whole. Specifically, if there is some redundancy of information (inter-correlation) among the $p$ original variables, then we may anticipate that a subset of (say) two or three principal axes will capture a substantial portion of the total variation in the system. A configuration plot of the positions of the points along the first two (or three) principal axes can therefore be examined in an attempt to visualise patterns among the sampling units along these major axes of variation.
To assess the utility of a 2-d (or 3-d) PCA/PCO plot, one might simply examine the percentage of the total variation that is extracted by the first 2 (or 3) axes. If this is substantial (e.g., 70% or 80%), then we may consider the plot to be useful for interpretation. Another useful tool is a scree plot, which shows the percentage of variation explained by sequentially ordered principal axes.
This does not, however, provide us with a meaningful measure of how well the plot (of reduced dimension) represents the (high-dimensional) inter-point distances (or dissimilarities) in the original multivariate space.
A good way to do that would be to calculate the stress associated with the 2-d (or 3-d) configuration. The notion of stress is already quite familiar as a measure of the adequacy of an ordination plot produced using multi-dimensional scaling (MDS, see section 5.2 and section 5.8 in Change in Marine Communities, 3rd edition). Indeed, it is the specific goal of the MDS algorithm to create a configuration that minimises stress. Of course, neither PCA nor PCO are specifically designed to minimise stress, so a PCA or PCO configuration will always (necessarily) have higher stress than an MDS configuration for a given data set. Nevertheless, having a measure of stress for a PCA/PCO configuration can help us assess its utility for displaying inter-point relationships faithfully. Here, we would consider that the usual rule-of-thumb of stress < 0.20 indicates a usefully interpretable configuration. Also, a plot of the configuration distances vs the original inter-point distances - a Shepard diagram - is clearly also a useful diagnostic tool here.
Scree Plot, Shepard diagram and Stress
In the PCO routine (accessed from a resemblance matrix by clicking on PERMANOVA+ > PCO...), PRIMER 8 now offers you new options to output these diagnostics tools:
- a Scree plot;
- Shepard diagrams in 2D and/or 3D; and
- Stress calculated using either metric MDS or threshold metric MDS
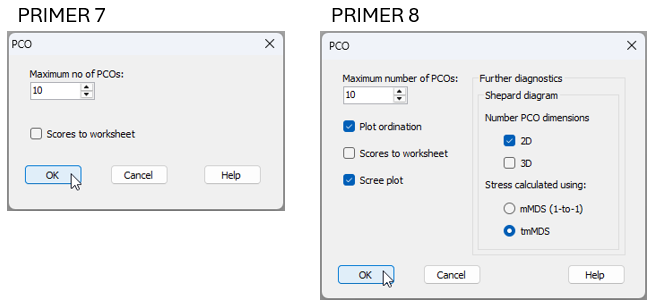
Fig. 15.8 compares the dialog window for the PCO routine in PRIMER 7 versus PRIMER 8.
Fig. 15.8 Comparison of the PERMANOVA+ > PCO... dialog window in PRIMER 7 (at left) vs PRIMER 8 (at right).
These same new diagnostic output options are also available for the PCA routine in PRIMER 8 (obtained from a data matrix by clicking on Analyse > PCA...).
It is important to recognise that these additional diagnostics do not in any way change anything about the fundamental PCA or PCO analysis itself that is being done. The calculation of principal axes for PCA or PCO ordination still remains exactly the same as ever, and may best be thought of as a projection. What is new and different here is simply the opportunity also to view the stress and the Shepard diagrams which attend the resulting PCA or PCO configuration. These diagnostics also help put these methods into perspective vis-à-vis any MDS ordination solutions one might obtain for a given dataset.
Example: Messolongi lagoon diatoms
Let's consider a study of diatom assemblages (densities of 193 species) at 17 sites in the lagoons of Messolongi, Aitoliko and Kleissova in Eastern Central Greece ( Danielidis (1991) ). Data from this study are contained in the file called 'Messolongi_diatom_density.pri', located in the 'Examples_P8' > 'Messolongi_diatoms' folder.
In what follows, we shall create an ordination of these data on the basis of a Bray-Curtis resemblance matrix calculated from square-root transformed densities, using each of the following methods:
- Non-metric MDS (nMDS)
- Threshold-metric MDS (tmMDS)
- Metric MDS (mMDS); and
- Principal co-ordinates analysis (PCO).
We will compare and contrast not only the ordination diagrams produced, but also the associated diagnostic Shepard diagrams and the values of stress. For PCO, we can produce Shepard diagrams and stress values in two different ways: (i) for comparison with tmMDS; or (ii) for comparison with mMDS.¶
- Open up the data matrix (Messolongi_diatom_density.pri) in PRIMER 8.
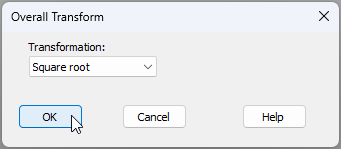
- From the data matrix (Messolongi_diatom_density), click Pre-treatment > Transform(overall)... and choose Square-root.
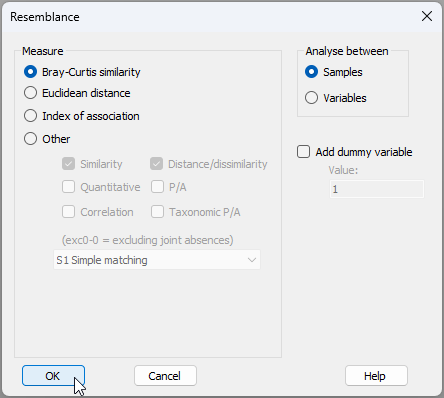
- The square-root transformed data are held in the sheet named 'Data1'. From this, click Analyse > Resemblance... and choose to calculate Bray-Curtis resemblances among samples, like so:
Non-metric MDS (nMDS)
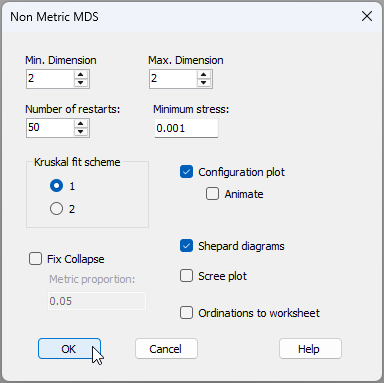
- The Bray-Curtis similarities are held in the sheet called 'Resem1'. From this, click Analyse > MDS > Non-metric MDS (nMDS).... Change the 'Max. Dimension' from 3 to 2 (for this exercise, we shall focus only on the 2-d solution), then click 'OK', as shown in the dialog below.
In the results ('MultiPlot1'), we can see the nMDS ordination ('Graph1'), the monotonic relationship shown in the Shepard diagram ('Graph2'), and that the value of stress is 0.092.
Threshold metric MDS (tmMDS)
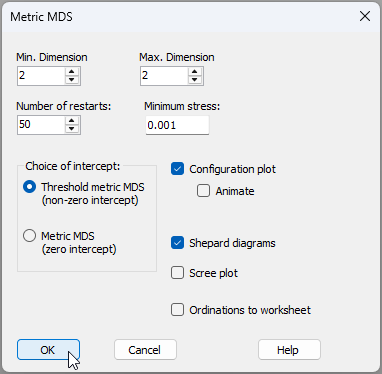
- From 'Resem1', click Analyse > MDS > Metric MDS (mMDS / tmMDS)..., then choose 'Choice of intercept: $\bullet$Threshold metric MDS (non-zero intercept)', change the 'Max. Dimension' from 3 to 2, then click 'OK' (see below).
In the results ('MultiPlot2'), we can see the tmMDS ordination ('Graph3'), the linear relationship shown in the Shepard diagram ('Graph4'), with a non-zero intercept of 60.54% similarity, and a stress value of 0.130. This is a bit larger than the stress obtained for the nMDS (0.092), but is still sufficiently low (< 0.20) to permit useful interpretations of the patterns of inter-point relationships seen in the diagram.
Note that the threshold (intercept) value of 60.54% similarity indicates that two points that are 'on top of one another' in the tmMDS configuration are to be interpreted as being ca. 39.46% dissimilar, and not 0% dissimilar.
Also, unlike the nMDS plot (which has no labels on its axes and from which only rank-order relationships among the points can be interpreted), the axes are labeled on the tmMDS plot, with values given in the units of the original (Bray-Curtis) resemblance measure. Thus, two points that are (say) 20 units apart from one another on this plot can be interpreted as being approximately (20% + 39.46%) = 49.46% dissimilar. Thus, although the tmMDS comes at the price of higher stress compared to the nMDS, it also 'gives something back' in the form of added interpretability (i.e., with labels on the axes permitting the estimation of actual inter-point dissimilarities). So, clearly, tmMDS can be a good option, provided the stress still remains low enough to yield a plot that is adequate for interpretation.
Metric MDS (mMDS)
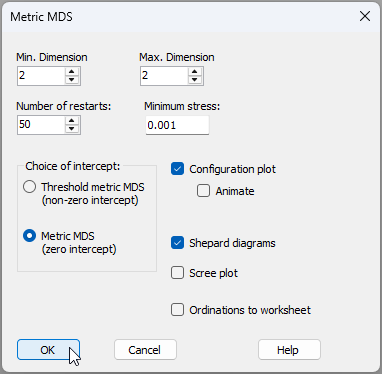
- From 'Resem1', click Analyse > MDS > Metric MDS (mMDS / tmMDS)..., then choose 'Choice of intercept: $\bullet$Metric MDS (zero intercept)', change the 'Max. Dimension' from 3 to 2, then click 'OK', viz:
In the results ('MultiPlot3'), we can see the mMDS ordination ('Graph5'), the linear relationship with a zero intercept in the Shepard diagram ('Graph6'), and a stress value of 0.252. This is much larger than that of either the nMDS (0.092) or the tmMDS (0.130), and is so high (> 0.25) as to prevent meaningful interpretation of any patterns seen in the resulting diagram.
Principal Co-ordinate Analysis (PCO)
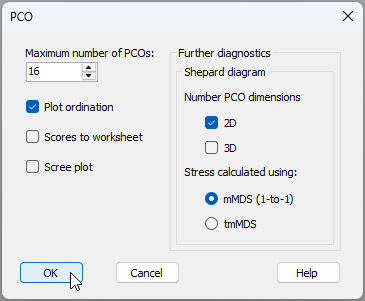
- First, we will run the PCO and get diagnostics for a Shepard diagram and associated stress calculation that uses a zero intercept (as in mMDS). From 'Resem1', click PERMANOVA+ > PCO, then choose 'Shepard diagram > Stress calculated using: $\bullet$mMDS (1-to-1)', then click 'OK' (see the dialog below).
The resulting PCO plot ('Graph7') and associated diagnostic Shepard diagram ('Graph8') are shown in 'MultiPlot4' in the Explorer tree.
The first two principal axes explain 44.56% of the total variation (less than half, as seen in the output file called 'PCO1' in the Explorer tree). In addition, the stress is extremely high, at 0.30, indicating that many of the inter-point distances seen in the PCO plot do not reflect their true dissimilarity well at all. This is especially true for the small to middle-sized dissimilarities that range from about 40% to 70% (see the large scatter of points in 'Graph8' for values having ~ 60% to 30% similarity along the x-axis).
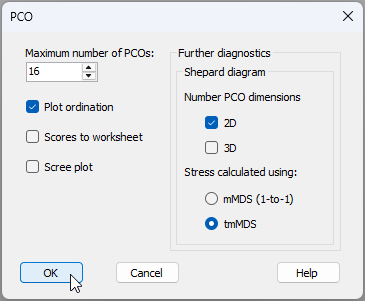
- Next, we will run the PCO again, but this time choose 'Shepard diagram > Stress calculated using: $\bullet$tmMDS' (see the dialog below).
The results are given in 'MultiPlot5' in the Explorer tree.
It is clear that the PCO plot itself remains exactly the same (compare 'Graph9' with 'Graph7'). However, we are now using a different yardstick to measure the stress associated with this plot. Specifically, we are permitting the intercept to float away from the origin, and asserting (as we did with the tmMDS) that there is a threshold dissimilarity that a pair of samples must exceed before they will occupy two different positions on the configuration plot. Here, the intercept is a similarity of 63.24%, corresponding to a dissimilarity threshold of 36.76%. In other words, any two samples that appear on top of each other on the plot (at a distance of zero) can be estimated to be 63.24% similar, not 100% similar.
By adding this flexibility, we can see that the stress improves dramatically, dropping to 0.201, which borders on becoming acceptably interpretable, at least for the broader structures. Once again, it is the larger dissimilarities that are captured better by the PCO (see the tighter clustering of points around the line-of-best-fit in the upper right area of the Shepard diagram), compared to the smaller ones.
Observations & recommendations
This example serves to demonstrate a more general point. Namely, for a given dataset, the ordering of methods with respect to the value of stress associated with the final configuration will almost always be: nMDS < tmMDS < mMDS < PCO. This is so because:
- all MDS solutions will, by their very design, achieve lower stress than a projection-type ordination, such as PCO, as they are explicitly geared to minimise stress; and
- within the MDS family of methods (nMDS, tmMDS and mMDS), nMDS has the fewest constraints, requiring only a monotonic relationship, while mMDS has the greatest constraints, requiring not just a linear relationship but also a zero-intercept in the Shepard diagram. This is more difficult to achieve, resulting in a higher stress.
Our general recommendations for ordination are:
- Use nMDS, as starting point, to achieve the lowest-stress solution.
- If the Shepard diagram for the nMDS shows that the relationship between configuration distances and original distances is approximately linear, then tmMDS can be used to achieve an ordination with the added benefit of having axes which (along with the added threshold dissimilarity) can be used to estimate actual inter-point dissimilarities directly from the plot. Only retain the tmMDS, however, if the stress is sufficiently low to permit interpretability ( < 0.20 as a rule-of-thumb).
- Only go one step further to consider using mMDS if the intercept is sufficiently close to zero to warrant this. (This will be evident in the Shepard diagram).
- PCO will typically not be the best tool to use, in general, to visualise inter-point relationships in an ordination, but will extract the axis (or axes) of greatest total variation through the data cloud as a whole. This means that large dissimilarities will typically be represented much better than small dissimilarities in a PCO plot.
Utility of PCO
Although PCO may not be the tool of choice for ordination, it is important to recognise that it is still a very useful tool for other reasons. Specifically, a variety of PERMANOVA+ routines use PCO ('under the hood') in order to calculate correctly (using the full set of PCO axes, and carefully accounting for negative eigenvalues):
- Distances among centroids (in the space of a chosen resemblance measure);
- Distances from individual points to their own group centroid (in PERMDISP); and
- Monte Carlo p-values (in PERMANOVA).
PCO axes are also used (obviously!) to perform canonical analysis of principal co-ordinates (CAP).
¶It is sometimes stated that PCO and metric MDS are the same thing. This is simply incorrect and the example here clearly demonstrates this. PCO is a projection onto principal axes, whereas metric MDS is distance-preserving and minimises stress for a configuration drawn in a chosen number of dimensions. PCO may sometimes be referred to as 'classical scaling', but it should not be confused with multi-dimensional scaling (MDS), metric or otherwise.

![17.Messolongi_data_only[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/17-messolongi-data-only-i.png)
![21.nMDS_results[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/21-nmds-results-i.png)
![23.tmMDS_results[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/23-tmmds-results-i.png)
![25.mMDS_results[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/25-mmds-results-i.png)
![27.PCO_w_mMDS_Shepard_results[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/27-pco-w-mmds-shepard-results-i.png)
![29.PCO_w_tmMDS_Shepard_results[i].png](https://learninghub.primer-e.com/uploads/images/gallery/2025-12/scaled-1680-/29-pco-w-tmmds-shepard-results-i.png)