6.7 Heterogeneity in more complex designs
Handling heterogeneity with multiple factors
The most important question to answer when you are dealing with a multi-factor study design and you decide you want to account for heterogeneity in dispersions (if present) is to answer the following question: Wherein does heterogeneity lie? Once you know which factor groupings (or which cells corresponding to combinations of factors) in the study design actually define the groups of sampling units that have heterogeneous dispersions (if any), then you can articulate this precisely in the PERMANOVA dialog and you are good to go.
The modification from $F_1$ to $F_2$ is readily extended to accommodate tests of individual terms in more complex (PERM)ANOVA designs. This is so because the PERMANOVA routine in PRIMER always constructs the pseudo $F$ statistic in such a way that the numerator and denominator have the same expectation under a true null hypothesis. Both $F_1$ and $F_2$ share this property, with the latter accounting for heterogeneity.
PERMANOVA, as implemented in PRIMER, will construct the correct test for every individual term in any given study design, by careful reference to the expectations of mean squares. To get the right test in every case, expectations of mean squares are used not only to construct the correct $F$ ratio, but also to discern the correct reduced-model residuals and the appropriate permutable units to permute. Every term will require its own denominator and its own permutation algorithm, which depends on whether terms are fixed or random or finite, whether there are nested terms, covariates, interactions, etc.
Futhermore, for unbalanced cases (which are of special interest to us here, of course), the 'Type' of sum of squares is also very important for the partitioning, the expectations of mean squares and subsequent tests. All of this is true whether you use $F_1$ or $F_2$, and it is very reassuring to know that PERMANOVA will do the right thing, precisely in accordance with your choices and your specific study design. No other software that we know of accomplishes all of this.¶
Wherein does heterogeneity lie?
In a multi-factor study design, it will be important to identify precisely where in the model heterogeneity (if any) might lie.† The most natural starting point will be to consider the cells that correspond to all combinations of the factors as your 'groups' (i.e., as if 'cells' were identified as a single factor in a one-way model). You can run a PERMDISP to compare dispersions among these cells, thus examining the null hypothesis of homogeneity in the dispersions of residuals, then go from there.
Nested design
Suppose you had two factors in a nested design (say, factor A = Locations and factor B = Sites nested within Locations), then the sources of variation in the model would be:
- A
- B(A)
- Residual
In this case, it is possible that the dispersion of replicates within each site differ among the sites. But it is also possible that the dispersion of the site centroids within each location might differ among locations. To examine each of these possiblities, in turn, you would need to do the following:
- (1) Test H01: dispersions of replicates within sites are equal across all sites.
- Do a PERMDISP test to compare dispersions among 'Sites' (ensuring that each site is labeled uniquely across the entire study design);
- (2) Test H02: dispersions of site centroids within locations are equal across all locations.
- Create a matrix of dissimilarities among the site centroids (using PERMANOVA+ > Distance Among Centroids... in PRIMER for this task), then
- From the resulting resemblance matrix among all sites, do a PERMDISP test to compare dispersions among 'Locations'.
Depending on the outcome from these tests, you could then decide whether you needed to use $F_2$ and, if so, which term in the model identifies the groups that have different dispersions. If (1) is significant, then 'Sites(Locations)' identifies heterogeneous groups, but if (2) is significant, then 'Locations' identifies heterogeneous groups. It is possible that neither of these PERMDISP tests come out as significant, in which case you can just use $F_1$. However, if both PERMDISP tests come out as significant, then you will need to consider running PERMANOVA twice (using $F_2$), first accounting for heterogeneity among the sites (to test B(A) correctly), then accounting for heterogeneity among the locations (to test A correctly). This will permit you to test each term in the model in a way that accounts for the heterogeneity present at each of these two different levels (i.e., two different scales of spatial variability) inherent in your study design.†
Crossed design
Suppose that you had two factors in a crossed design (say, factor A = Treatments and factor B = Locations, crossed with Treatments), then the sources of variation in the model would be:
- A
- B
- A $\times$ B
- Residual
Just as before, we will want to begin by discovering wherein heterogeneity (if any) might lie. First, we need to test the null hypothesis of homogeneity among the cells. We would proceed as follows:
- Create a factor corresponding to all combinations of factors A and B (e.g., using Edit > Factors... > Combine... in PRIMER). We might call this new factor 'AB'.
- (1) Test H01: dispersions of replicates within AB cells are equal across all the cells.
- Do a PERMDISP test to compare dispersions among levels of the newly created factor 'AB'.
If the test in (1) above is statistically significant, we may then proceed to perform a PERMANOVA using $F_2$ and identify the interaction term 'A $\times$ B' as being the one that identifies the groups ('cells') with heterogeneous dispersions.
If the test in (1) above is not statistically significant, then before considering heterogeneity among groups associated with either of the main effects, we need to first do a PERMANOVA to investigate the possibility that the two factors interact with one another. Our next step is therefore:
- (2) Test H02: there is no interaction between factors A and B in their effects on the centroids.
- Do a two-way crossed PERMANOVA with factors 'A' and 'B' and specifically examine the test of the term 'A $\times$ B'.
Why do we need to do this test (2) above? Well, it is because...
Interactions can generate patterns of heterogeneity in main effects
It is useful at this juncture to point out why a PERMDISP test done on either of the main effects alone, ignoring the other factor, might be misleading. In essence, if two factors interact with one another (in a PERMANOVA), then this could (unhelpfully) be detected as 'heterogeneity' in one or other of the main effects.
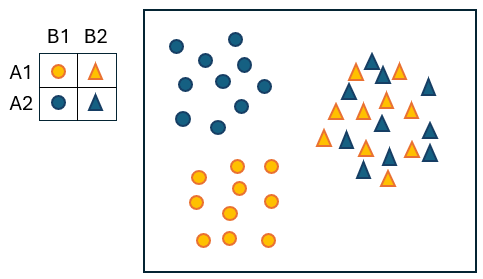
To see how this can happen, suppose there are $a = 2$ treatments and $b = 2$ locations, and suppose also that these two factors interact. More specifically, suppose the interaction is caused by there being significant effects of factor A ('Treatments') on the centroids at one of the locations ('B1'), but not at the other ('B2'), as shown in Fig. 6.3.
Fig. 6.3. Schematic diagram of a two-way crossed design where there is an interaction between factor A (with levels: A1 = blue and A2 = orange) and factor B (with levels: B1 = circles and B2 = triangles), demonstrating how interpreting the results of a test for differences in 'dispersion' of a main effect (e.g., factor B here) can be confounded by interactive effects of another factor (factor A here) on the centroids.
Clearly, if we did a PERMDISP test to compare the dispersions of the 2 groups corresponding to factor B (i.e., circles vs triangles), the result would be statistically significant. The graphic (Fig. 6.3) clearly shows that if you ignore factor A (i.e., if you ignore the colour of the symbols), then the circles are more spread out (dispersed) than the triangles. However, this actually has nothing to do with differences in 'dispersion' at all (all 4 of the A $\times$ B cells have roughly equal spread), but rather is due to the significant interaction in centroid effects. Specifically, at location B1 (circles) we see a shift in centroid due to factor A (A1 $\ne$ A2), but at location B2 (triangles), we do not see a shift in centroid due to factor A (A1 $\approx$ A2).
Continuing onwards now with our logical flow... If the test of (2) above is significant, then we effectively proceed with interpreting all of the results given in the PERMANOVA, including possibly doing relevant pairwise comparisons and associated ordination plots, etc.
If the test in (2) above is not significant, then we might consider doing the following two tests:
- (3) Test H03: dispersions of replicates within groups defined by factor A (ignoring factor B) are equal.
- Do a PERMDISP test to compare dispersions among levels of factor 'A'.
- (4) Test H04: dispersions of replicates within groups defined by factor B (ignoring factor A) are equal.
- Do a PERMDISP test to compare dispersions among levels of factor 'B'.
Depending on the outcomes of these tests (3) and (4), you could then do the two-factor PERMANOVA and account for heterogeneous dispersions in either of these factors, if needed.†
A few take-home messages
Handling potential heterogeneity of dispersions in multi-factor (PERM)ANOVA designs can potentially become very complex, as we have seen even in the two-way cases outlined above. However, it is important not to get too de-railed from the main game of your study, and there is no need to feel overwhelmed by this topic. Let's summarise a few take-home messages about all of this:
- PERMANOVA is very robust to heterogeneity if your design is balanced. If sample sizes are equal, you can use $F_1$ in the usual way for PERMANOVA and rest assured that the tests for centroid differences, interactions, etc. for all terms in the model generally will be quite robust and interpretable, regardless of any heterogeneity.
- If your design is unbalanced, test for heterogeneity in the highest-order cells and accommodate it. A useful standard approach for an unbalanced multi-factor design will always be to start by performing a PERMDISP on the highest-order cells in your study design. In other words, if you have 3 factors (A, B, and C), then create a factor that corresponds to all combinations of levels of those factors (A $\times$ B $\times$ C) and do a PERMDISP on that new 'combined' factor. If heterogeneity is present (among those cells), you can then easily accommodate it in your PERMANOVA by ticking the box to use $F_2$ ('Allow for heterogeneity') and nominating the highest order interaction term (e.g., A $\times$ B $\times$ C) in the PERMANOVA design file as the 'Term identifying groups with different dispersions'.
- Take one step at a time and think logically about each step in your testing procedure. If there is no heterogeneity in the dispersions of replicates among the highest-order cells in your study design, then you can proceed to examine a suite of logical hypotheses regarding differences in centroids and/or dispersions associated with other terms in your model. Usually, you would start with the higher-order terms in the model (towards the bottom of a PERMANOVA table of results) and gradually 'work your way up' towards considering the main effects. This might take some time and care, depending on the design and the number of simultaneous factors you are dealing with. Often, a helpful thing to think about is how the sum of squares (SS) for each term itself is constructed. This will point you to thinking about the right way to construct a test for homogeneity for any given factor or term in the model.‡ You will also have to consider how potential interactions (in centroid effects) among factors might alter your perception of dispersion differences across the main effects.
- If you have a (modestly) unbalanced design with (modest) heterogeneity, PERMANOVA is still quite a robust test. PERMANOVA is definitely focused on the null hypothesis of no differences in centroids. It is unlikely that modest differences in dispersion here or there are going to adversely affect your inferences drawn broadly from PERMANOVA tests much at all, especially if the degree of imbalance in your sample sizes is not dramatic (e.g., if you just have the odd replicate missing here or there in a few cells). Sometimes, the added complexity of dealing with heterogeneity (particularly if it occurs at multiple different levels and/or for more than one factor in your study design) can outweigh the benefits of attempting to accommodate it.
- Bear in mind that it takes quite a few replicates even to measure and compare dispersions in the first place. If you have very small sample sizes per cell (e.g., less than 4 or 5), then formal statistical comparisons of cell dispersions using PERMDISP are probably not worth much (i.e., they can be somewhat unreliable). Even in univariate analysis, you need far more replicates to get a decent estimate of the variance of a population than you would need to have in order to get a good estimate of the population mean. This is true also for multivariate dissimliarity-based analyses. So, if you have small sample sizes per cell, proceeding with the 'vanilla-flavoured' PERMANOVA (using $F_1$) to compare centroids (as you do not have much information even to estimate the dispersions for the cells) is quite a reasonable course of action.§
- Be cautious if you wish. If the design is unbalanced and the replication per cell is low, you can alternatively choose to take a more conservative stance: simply assume that there is heterogeneity among the cells, and do a PERMANOVA using $F_2$ accordingly. Taking that approach would be defensible, but perhaps would lack power.
¶ Note that adonis2 in the vegan package in R will not do any of this. Please see our recent exposé on this topic. In addition, no other software package that we know of (in R or otherwise) will implement PERMANOVA using $F_2$ or in any other way that accounts correctly for heterogeneity.
† In PRIMER 8 you can currently only specify one source of heterogeneity at a time in any given PERMANOVA model, although theoretically the general approach we use with $F_2$ need not be restricted necessarily in this way. It is only a matter of working out how to logistically accommodate multiple sources of heterogeneity simultaneously. Although this problem is not trivial, it is solvable. If you do have multiple sources of heterogeneity, then you will need potentially to consider running PERMANOVA more than once to account for this in the right way for tests of different terms in the model.
‡ For example, the SS for 'Locations' in the Nested design discussed on this page is constructed as the sum of squared deviations of 'Site' centroids around the 'Location' centroids. So that points us to: (i) first get the dissimilarities among the Site centroids, then (ii) do a PERMDISP for the location factor using those Site centroids as the 'replicates'.
§ Small sample sizes in cells may well have other consequences for your inferences, of course, such as limitations on the numbers of possible permutations for pairwise comparisons, thus limiting the precision of resulting p-values and, hence, low power.